Mahdi Yousefian
Iran
Harnessing Base editing with CRISPR-Cas13 and machine learning for precision gene therapy in rare genetic disorder
Mahdi Yousefian1, Maryam Baharmast2
1Islamic Azad University of Tehran Medical Science , www.Orcid.org/0009-0006-7619-523X, www.Linkedin.com/in/mahdiyousefian ,Yousefian.Mahdi2004@gmail.com
2Islamic Azad University of Tehran Medical Science , www.Orcid.org/0009-0001-0839-9779,, www.Linkedin.com/in/maryam-baharmast
Abstract
Background
Sickle cell disease (SCD) is a genetic disorder caused by the inheritance of abnormal beta-globin alleles carrying the sickle mutation (Glu6Val, βS) in the HBB gene, leading to the characteristic crescent-shaped, or “sickleshaped,” erythrocytes and impaired oxygen transport. This disease results in vaso-occlusion, repeated ischemia, and inflammation, associated with various cellular and plasma factors and abnormal endothelial interactions, causing severe acute and chronic pain, immunodeficiency, multi-organ failure, and early death.
To date, several clinical treatments have been applied; however, access to these advanced therapies is often limited by cost and donor availability. The only curative modality remains stem cell transplantation .Gene therapy, however, is currently undergoing rapid clinical investigation as a potential cure for SCD.
Recently, a revolutionary tool for RNA editing has emerged: the CRISPR–Cas13 system, which shows high potential for advancing nucleic acid therapeutics. Unlike the DNA-targeting CRISPR–Cas9 system, Cas13 targets and cleaves RNA, enabling gene silencing without inducing genomic instability. This approach can suppress disease-causing genes, correct splicing errors, and modulate immune responses. Despite these advances, challenges remain, including the need to refine targeting specificity, mitigate off-target effects, and ensure effective in vivo delivery.
This review provides an overview of the characteristics and mechanisms of the CRISPR–Cas13 system, as well as the recent enhancements achieved through machine learning technologies.
Methods
Currently, Cas13 is being applied for transcript knockdown, RNA base editing, and live-cell applications. It has been determined that CRISPR-Cas13 achieves target specificity via a 28–30-nt spacer and requires a guide RNA of approximately 64 nt. Uniquely, Cas13 exhibits collateral activity post-recognition, leading to non-specific degradation of surrounding transcripts. Although this phenomenon occurs in bacteria, it does not manifest in plant or mammalian cells, enabling its potential development as an RNA-targeting tool.
One promising application is its use in diagnostics, where Cas13 can be engineered to specifically and accurately cleave fluorescent reporters upon target recognition. Cas13a and Cas13b have demonstrated efficacy as RNA knockdown tools, while Cas13d orthologs can control endogenous transcript splicing. Notably, Cas13d’s small size (~930 aa), the smallest among class 2 CRISPR effectors, facilitates its potential in vivo delivery.
Results
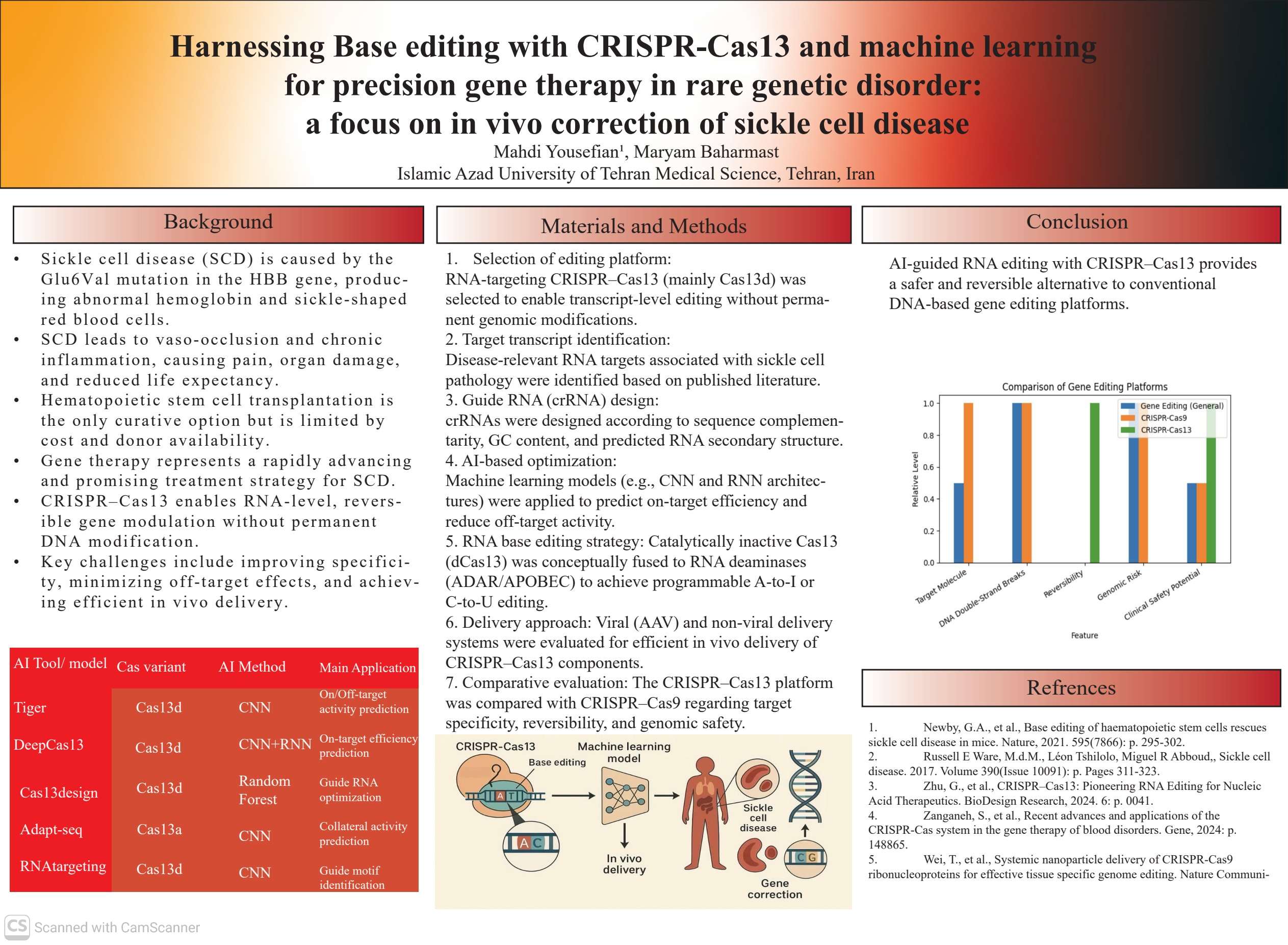
This study highlights the transformative potential of Cas13-based base editing technologies, combined with machine learning, to achieve precise and efficient RNA editing for therapeutic applications such as sickle cell disease (SCD). While traditional CRISPR-Cas9 genome editing remains a powerful tool for correcting pathogenic mutations, it is inherently limited by risks such as off-target genomic alterations, double-stranded breaks, and potential tumorigenesis. Our review demonstrates that integrating AI models with Cas13 base editing platforms significantly enhances target specificity and editing efficiency. Machine learning models such as TIGER and DeepCas13 enable accurate prediction of guide RNA efficacy by analyzing complex features including nucleotide composition, RNA secondary structures, and target accessibility.
Conclusions
Remarkable progress has been achieved over the last decade in studying and manipulating genes using CRISPR-Cas systems. CRISPR-Cas13 systems, in particular, provide valuable means for making transient modifications, offering unique opportunities for therapeutic applications. The integration of AI with genome editing has enabled personalized treatments by optimizing guide designs and minimizing off-target effects.
Focusing on the in vivo correction of sickle cell disease, programmable RNA editors guided by predictive computational models have demonstrated enhanced on-target accuracy with minimal off-target activity. Despite current limitations in in vivo applications, CRISPR-Cas13 shows great promise for RNA base editing by avoiding permanent genomic alterations and mitigating unintended consequences. Real-time AI-driven tools in CRISPR applications hold significant promise for treating a wide range of RNA-mediated diseases and genetic disorders.

Leave A Comment